We develop artificial intelligence based methods for protein engineering and sampling of protein dynamics. You can find all our code on Shukla group GitHub page.
Artificial Intelligence Based Methods
EZspecificity: Enzyme specificity prediction using cross attention graph neural networks
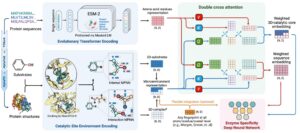
Enzyme specificity, a hallmark of biological catalysis, depends on the subtle three-dimensional complementarity between active sites and substrates. EZSpecificity leverages a cross-attention-empowered SE(3)-equivariant graph neural network to learn this structural logic, enabling accurate prediction of enzyme–substrate pairs from sequence and structure information. Trained on a comprehensive enzyme–substrate interaction database and validated experimentally, EZSpecificity achieved unprecedented precision in identifying reactive substrates, providing a universal framework for understanding and engineering enzymatic function across biology, chemistry, and medicine.
Haiyang Cui†, Yufeng Su†, Tanner J. Dean†, Tianhao Yu, Zhengyi Zhang, Jian Peng, Diwakar Shukla*, and Huimin Zhao*, Enzyme specificity prediction using cross attention graph neural networks. Nature (2025). Publication Link | Demo Link
ESMDynamic: Fast and Accurate Prediction of Protein Dynamic Contact Maps from Single Sequences
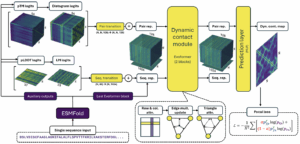
ESMDynamic is a deep learning model that predicts dynamic residue-residue contact probability maps directly from protein sequence. Built on the ESMFold architecture, ESMDynamic is trained on contact fluctuations from experimental structure ensembles and molecular dynamics (MD) simulations, enabling it to capture diverse modes of structural variability without requiring multiple sequence alignments.
Diego E. Kleiman, Jiangyan Feng, Zhengyuan Xue and Diwakar Shukla*, ESMDynamic: Fast and Accurate Prediction of Protein Dynamic Contact Maps from Single Sequences. bioRxiv, doi: 10.1101/2025.08.20.671365v1, 2025. Publication Link
An open-source implementation of ESMDynamic is available at https://github.com/ShuklaGroup/ESMDynamic
Multi-Objective Controlled Extrapolation for Protein Sequence Generation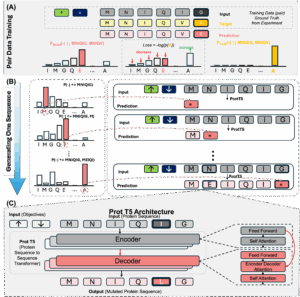
This work introduces a framework for multi-objective protein design that leverages pair-wise sequence information to create novel sequences outside the training distribution. By extending previous extrapolation frameworks to multiple objectives, we provide an effective method for designing protein sequences optimized for selectivity.
Brenda M. Wang, Nicole Chiang, Holly M. Ekas, Dylan M. Brown, Garrett Dildine, Tyler J. Lucci, Siyuan Feng, Vanessa Bly, Jean-François Gaillard, Julius B. Lucks, Ashty S. Karim, Diwakar Shukla, Michael C. Jewett*, Active learning-guided optimization of cell-free biosensors for lead testing in drinking water. bioRxiv, doi:10.1101/2025.08.20.671382, 2025. Publication Link
An open-source implementation of Multi-objective Controlled Extrapolation is available at https://github.com/ShuklaGroup/multiobjective_controlled_extrapolation
LassoESM: A tailored large language model for Lasso peptides
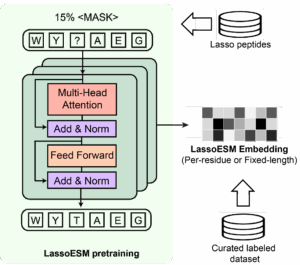 We developed a lasso peptide-specific language model (LassoESM) by leveraging advances in pre-trained PLMs to aid the prediction of lasso peptide related properties.
We developed a lasso peptide-specific language model (LassoESM) by leveraging advances in pre-trained PLMs to aid the prediction of lasso peptide related properties.
Xuenan Mi, Susanna L. Barrett, Douglas A. Mitchell* and Diwakar Shukla*, LassoESM: A tailored language model to enhance lasso peptide property prediction. Nature Communications, Volume 16, Article number: 8545, 2025. Publication Link
An open-source implementation of LassoESM is available at https://github.com/ShuklaGroup/LassoESM
PeptideESM: A tailored large language model for peptides
 We developed a peptide-specific language model (PeptideESM) by leveraging advances in pre-trained PLMs to aid the prediction of peptide related properties.
We developed a peptide-specific language model (PeptideESM) by leveraging advances in pre-trained PLMs to aid the prediction of peptide related properties.
Joseph D. Clark, Xuenan Mi, Douglas A. Mitchell and Diwakar Shukla*, Substrate prediction for RiPP biosynthetic enzymes via masked language modeling and transfer learning. Digital Discovery, Volume 4, Pages 343-354, 2025. Publication Link
An open-source implementation of PeptidesESM is available at https://github.com/ShuklaGroup/LazBFDEF
TLmutation: Protein Variant Effect Predictor based on transfer learning
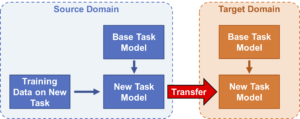 TLmutation leverages deep mutational scanning datasets to predict the functional consequences of mutations in homologous proteins.
TLmutation leverages deep mutational scanning datasets to predict the functional consequences of mutations in homologous proteins.
Zahra Shamsi, Matthew Chan and Diwakar Shukla*, TLmutation: Predicting the Effects of Mutations Using Transfer Learning. The Journal of Physical Chemistry B, Volume 124, Issue 19, Pages 3845-3854, 2020. Publication Link
An open-source implementation of TLmutation is available at https://github.com/ShuklaGroup/TLMutation
Molecular Dynamics Simulation Based Methods
Ensemble Adaptive Sampling Scheme (EASE)
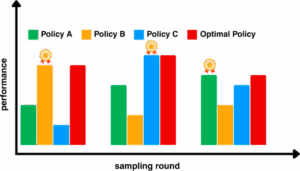
We developed a framework for identifying the optimal sampling policy through metric-driven ranking. Our approach systematically evaluates the policy ensemble and ranks the policies based on their ability to explore the conformational space effectively. This approach takes an ensemble of adaptive sampling policies and identifies the optimal policy for the next round based on current data.
Hassan Nadeem and Diwakar Shukla*, Ensemble Adaptive Sampling Scheme: Identifying an Optimal Sampling Strategy via Policy Ranking. Journal of Chemical Theory & Computation, Volume 21, Issue 9, Pages 4626–4639, 2025. Publication Link
An open-source implementation of policy ranking framework and EASE is available at https://github.com/ShuklaGroup/EASE.
Maximum Entropy VAMPNets (MaxEnt VAMPNets)
 Inspired by the active learning approach of uncertainty-based sampling, we also present MaxEnt VAMPNet. This technique consists of restarting simulations from the microstates that maximize the Shannon entropy of a VAMPNet trained to perform the soft discretization of metastable states.
Inspired by the active learning approach of uncertainty-based sampling, we also present MaxEnt VAMPNet. This technique consists of restarting simulations from the microstates that maximize the Shannon entropy of a VAMPNet trained to perform the soft discretization of metastable states.
Diego E. Kleiman and Diwakar Shukla*, Active Learning of the Conformational Ensemble of Proteins Using Maximum Entropy VAMPNets. Journal of Chemical Theory & Computation, Volume 19, Issue 14, Pages 4377–4388, 2023. Publication Link
An open-source implementation of MaxEnt VAMPNets is available on Github. https://github.com/ShuklaGroup/MaxEntVAMPNet.
Multiagent Reinforcement Learning-Based Adaptive Sampling (MA-REAP)

We have developed two algorithms inspired by multiagent RL that extend the functionality of closely related techniques (REAP and TSLC) to situations where the sampling can be accelerated by learning from different regions of the energy landscape through coordinated agents.
Diego E. Kleiman and Diwakar Shukla*, Multiagent Reinforcement Learning-Based Adaptive Sampling for Conformational Dynamics of Proteins. Journal of Chemical Theory & Computation, Volume 18, Issue 9, Pages 5422–5434, 2022. Publication Link
An open-source implementation of multi-agent REAP is available on Github. https://github.com/ShuklaGroup/MA_REAP
Reinforcement Learning-Based Adaptive Sampling (REAP)
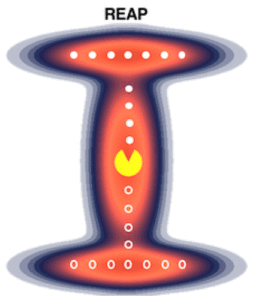 The REAP algorithm uses concepts from the field of reinforcement learning, a subset of machine learning, which rewards sampling along important degrees of freedom and disregards others that do not facilitate exploration or exploitation.
The REAP algorithm uses concepts from the field of reinforcement learning, a subset of machine learning, which rewards sampling along important degrees of freedom and disregards others that do not facilitate exploration or exploitation.
Zahra Shamsi, Kevin J. Cheng and Diwakar Shukla*, Reinforcement Learning Based Adaptive Sampling: REAPing Rewards by Exploring Protein Conformational Landscapes. Journal of Physical Chemistry B, Volume 122, Issue 35, Pages 8386–8395, 2018. Publication Link
An open-source implementation of REAP is available on Github. https://github.com/ShuklaGroup/REAP-ReinforcementLearningBasedAdaptiveSampling
Optimal Probes
 Optimal Probes is a software package to predict the smallest set of residue pairs for site-directed spin labeling for DEER/EPR experiments that best capture the slowest dynamics in a protein of interest. It uses molecular dynamics (MD) simulation datasets and a hyperparameter optimization strategy for a Markov state model (MSM).
Optimal Probes is a software package to predict the smallest set of residue pairs for site-directed spin labeling for DEER/EPR experiments that best capture the slowest dynamics in a protein of interest. It uses molecular dynamics (MD) simulation datasets and a hyperparameter optimization strategy for a Markov state model (MSM).
Shriyaa Mittal and Diwakar Shukla*. “Predicting Optimal DEER Label Positions to Study Protein Conformational Heterogeneity”. Journal of Physical Chemistry B, Volume 121, Issue 42, Pages 9761–9770, 2017.
An open-source implementation of Optimal Probes is available on Github. https://github.com/ShuklaGroup/optimalProbes/wiki
Evolutionary Coupling Guided Adaptive Sampling (ECAS)
In this method, we assess the use of distances between evolutionarily coupled residues as natural choices for reaction coordinates which can be incorporated into Markov state model-based adaptive sampling schemes and potentially used to predict not only functional conformations but also pathways of conformational change, protein folding, and protein-protein association.
Zahra Shamsi, Alexander S. Moffett and Diwakar Shukla*, Enhanced unbiased sampling of protein dynamics using evolutionary coupling information. Scientific Reports, volume 7, Article number: 12700, 2017.

